|
1/7/2024 0 Comments Protein backbone angles Therefore, the sampling-based feature and the efficiency of OPUS-Refine offer greater potentiality for it to take advantage of any other method to achieve better performance. Therefore, considerable effort is being put into developing computational methods for predicting backbone angles. In this work, we predict all of these four backbone angles using HMM profiles besides other informative features within a fully connected neural network (FCNN). However, reliable determination of backbone torsion angles using conventional experimental methods is slow and expensive. OPUS-Refine is a highly efficient framework, it takes only about 4 s to refine the torsion angles and 30 s to refine the global structure of a protein with 100 residues in length on a typical desktop personal computer. Protein backbone torsion angles ( and ) prediction is a key sub-problem in predicting protein structures. After refinement, compared to the predicted structures obtained from RaptorX online server, the accuracy of both global structural configurations (measured by TM-score and RMSD) and local structural configurations (measured by Phi/Psi) results are improved. Backbone angles and are measurements of the residue-wise torsion (Ramachandran et al., 1963), whereas angles and are spread over 3 (dihedral. Furthermore, we also introduce the contact map predicted by RaptorX to OPUS-Refine as a global structural constraint. Recognizing secondary structure as a coarse-grained description of protein backbone, more recent efforts have been shifted to the prediction of continuously valued backbone torsion angles. In addition, to facilitate the sampling efficiency, we construct a neighbor-dependent statistical torsion angles sampling database, namely, OPUS-TA, which may be useful for other sampling-based methods. Chemical shift measurements are less affected by peak overlap or reduced spectral sensitivity than measurements of peak. 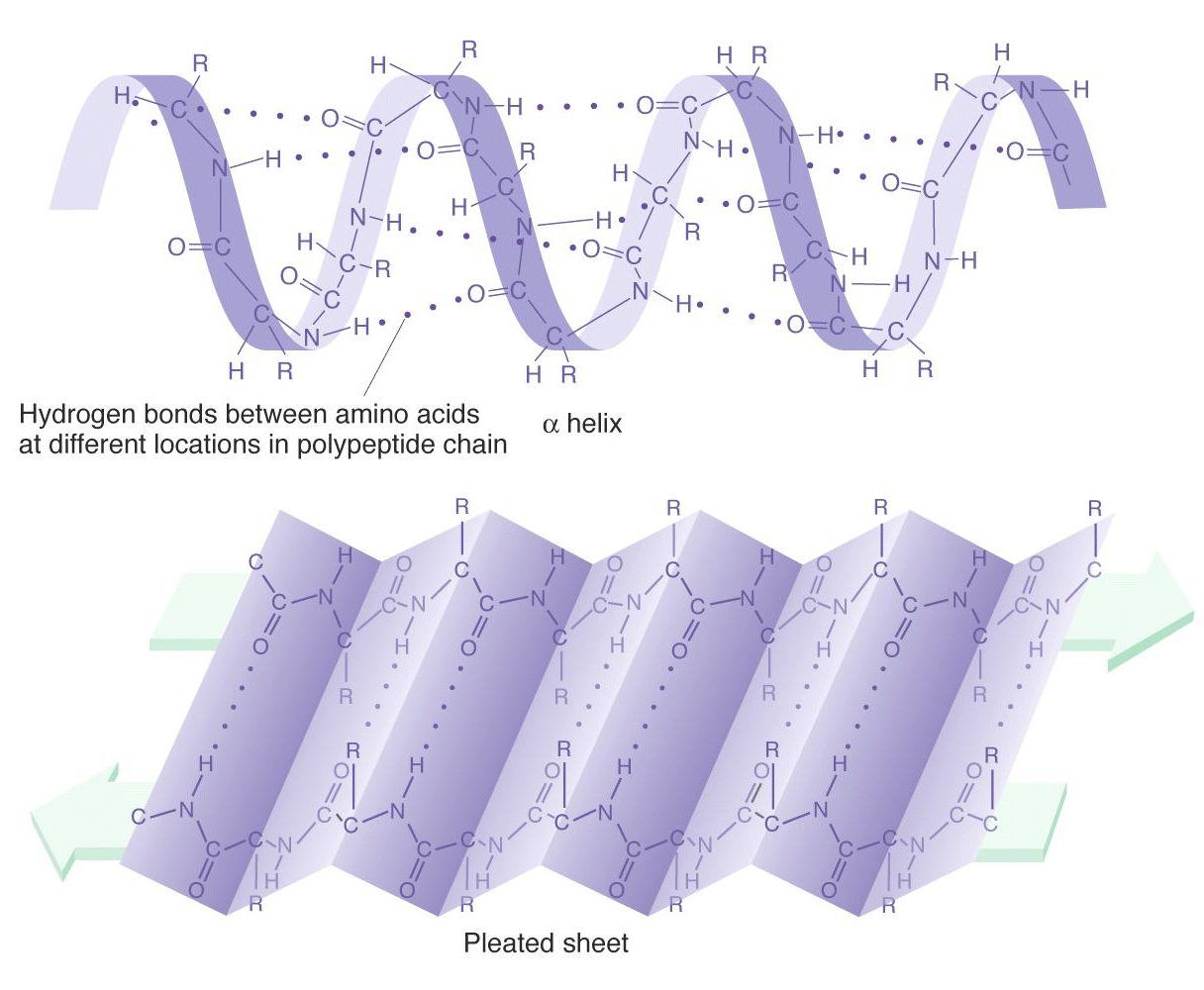
After OPUS-Refine refinement, for instance, the accuracy of Phi/Psi predicted by SPIDER3 and SPOT-1D are both increased. Indeed, it is well known that 1 H, 13 C and 13 CO shifts are very sensitive to backbone / angles, while 15 N shifts appear to be significantly influenced by side chain 1 angles of the preceding residue ( 11). OPUS-Refine is a sampling-based method, therefore, the results of other prediction methods can be used as its constraints. In this paper, we propose a general postprocessing method for all prediction methods, namely, OPUS-Refine, which may contribute to the field in a different way. Internal coordinates comprise Psi, Omega and Phi dihedral angles along the protein backbone, Chi angles along the sidechains, and all 3-atom angles and bond.  
Protein backbone torsion angles (Phi and Psi) are crucial for protein local conformation description. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
